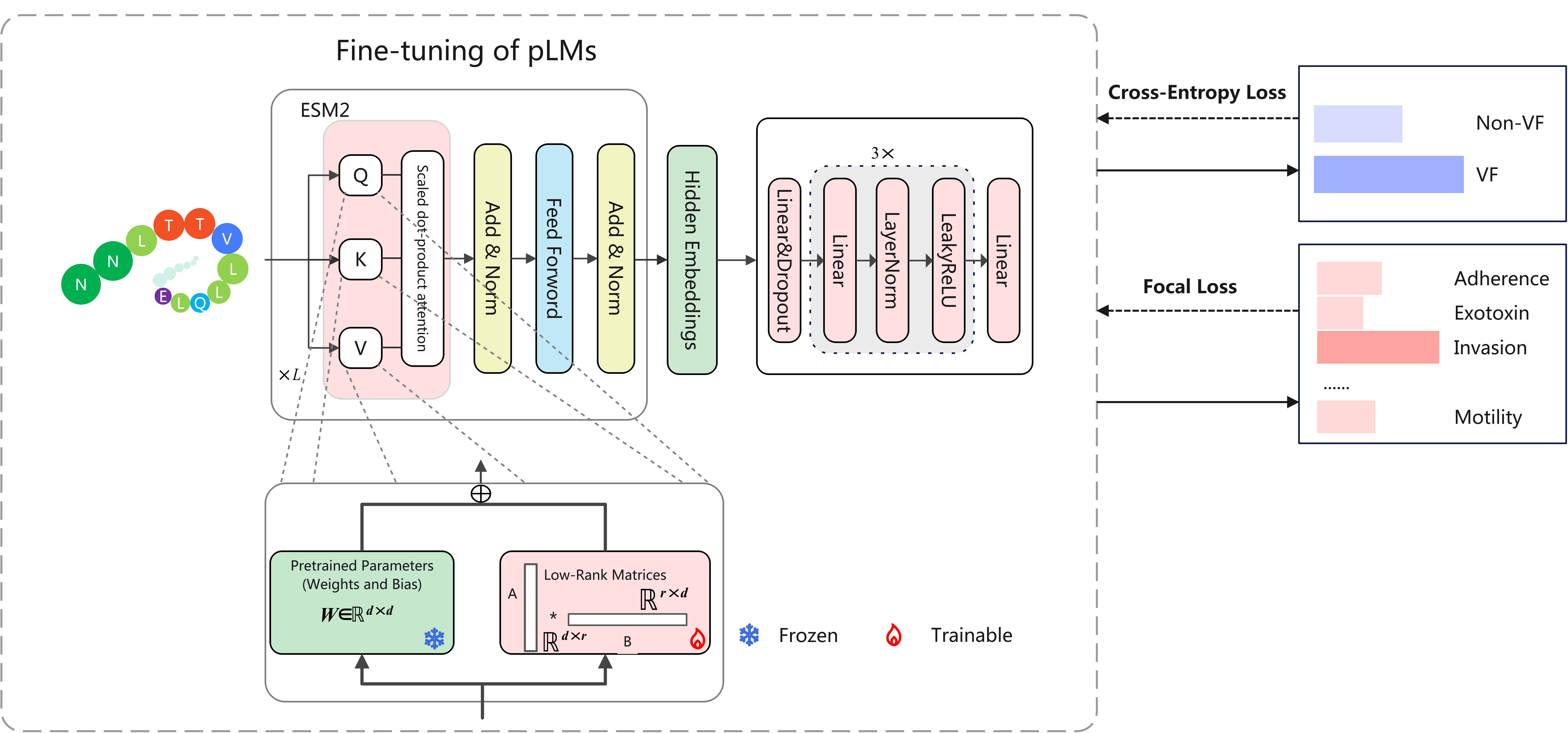
VirulentHunter is a novel deep learning framework designed to address the limitations of existing VF identification methods. Traditional methods primarily rely on homology alignment, which can miss novel or divergent VFs and lack effective means for VF functional classification. VirulentHunter works directly from protein sequences, using deep learning models to achieve simultaneous VF identification and classification. We have integrated multiple public resources to build a comprehensive and rigorously annotated VF database, providing a solid foundation for model training and prediction. The application of VirulentHunter will drive in-depth research on microbial pathogenicity, providing new perspectives for studying microbial pathogenicity in both controlled laboratory settings and complex environmental samples.
Instruction and Sample
Please note the following input requirements before filling in the form:
- Allowed file extensions: .faa, .fasta, .fastq, .fa, .fna
- Maximum file size for Protein file: 50MB
- Protein sequences are recommended for better transmission and computational efficiency.
- Sample File
Important Notice:
- Strain Genome and Metagenome analysis are temporarily unavailable due to limited computational resources.
- Processing time: Approximately 10 hours for 4,000 protein sequences (CPU-only, no GPU support).
- For faster processing, please consider local deployment of this tool.
Recent Projects Status
| Project ID | Submit Time(UTC) | Type | Status |
|---|---|---|---|
| raisonj****871f26ad | 2026-03-19 15:02 | protein | Pending |
| c.picho****65561c1a | 2026-03-19 09:05 | protein | Pending |
| bteben0****1ddb94cc | 2026-03-18 14:09 | protein | Pending |
| csungji****72d9c530 | 2026-03-18 08:02 | protein | Pending |
| prakash****d1f1df11 | 2026-03-18 04:29 | protein | Pending |
| ppvb30d****c70a0bac | 2026-03-17 19:20 | protein | Pending |
| pieter.****0b754080 | 2026-03-11 09:52 | protein | Pending |
| 1059496****f04ca874 | 2026-03-10 01:19 | protein | Pending |
| satyamv****87c238e8 | 2026-03-09 08:58 | protein | Pending |
| satyamv****1f84d17c | 2026-03-09 08:57 | protein | Pending |
| sean.na****036abbdb | 2026-03-05 07:33 | protein | Pending |
| aryacj1****884ddc7f | 2026-03-03 11:50 | protein | Pending |
| sayande****6ad5a066 | 2026-03-03 11:46 | protein | Pending |
| khnsp2m****1d7abb94 | 2026-03-03 10:20 | protein | Pending |
| maribas****2d6261ed | 2026-03-02 20:01 | protein | Pending |
| atiqabr****c06ead1c | 2026-02-25 05:48 | protein | Pending |
| per.kri****09d6890d | 2026-02-19 07:31 | protein | Pending |
| florenc****00a2f931 | 2026-02-18 18:41 | protein | Pending |
| nathani****77b46839 | 2026-02-18 16:52 | protein | Pending |
| shekhar****e798ed30 | 2026-02-18 04:32 | protein | Pending |
Tutorial
Here, we present VirulentHunter, a novel deep learning framework for simultaneous VF identification and classification directly from protein sequences. We constructed a comprehensive, curated VF database by integrating diverse public resources and rigorously expanding VF category annotations. Benchmarking demonstrates that VirulentHunter significantly outperforms existing methods, particularly for VFs lacking detectable homology.
Step 1: Navigate to the Search Tab
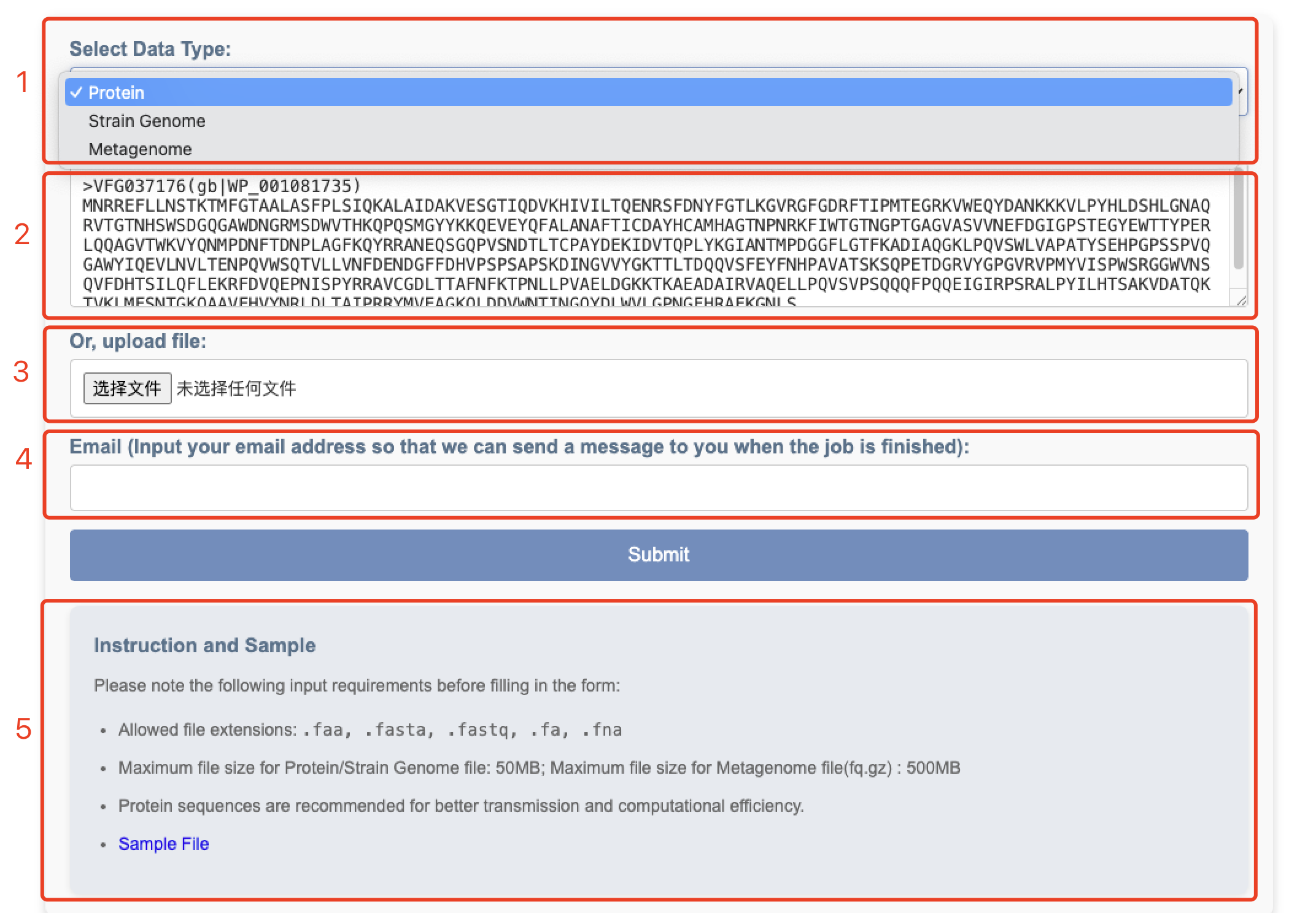
Upon clicking the Search tab, you will see the interface shown below:
1. Supported Input Data Types:
- Protein sequences
- Strain Genome
- Metagenome
2. Sequence Input Box:
For small datasets (protein sequences or bacterial strain genomes), directly paste
your sequences into this box. Note: This feature is unavailable for metagenomic data.
3. File Upload Box:
For larger datasets (bacterial strain genomes or metagenomic data), you can upload
your data in the FASTA format.
4. Email Input Box (Critical):
Enter your email address so that we can send you a message when the job is finished.
5. Additional Notes:
Brief reminders and instructions are displayed to guide your workflow.
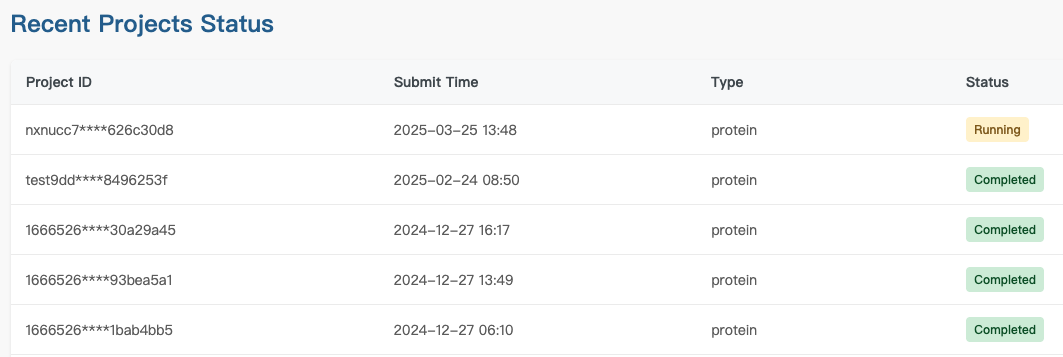
Step 2: Monitor Task Progress via the Status Tab
Click the Status tab to view the real-time progress of your tasks.
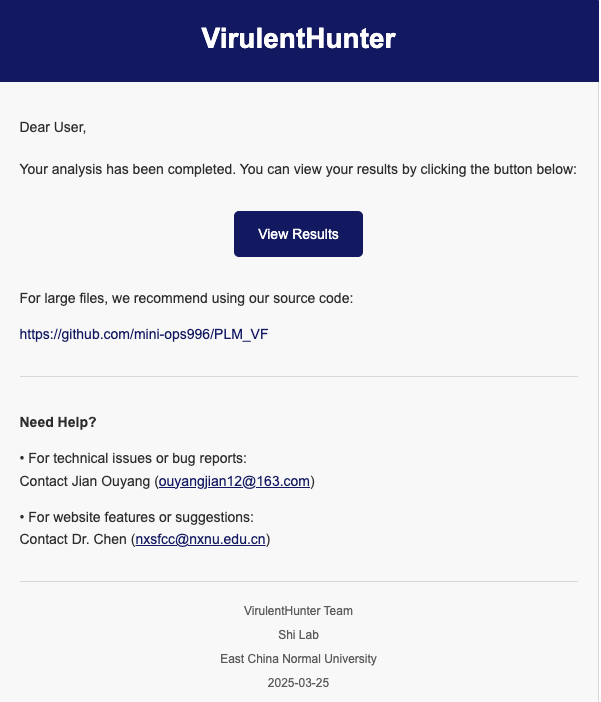
Step 3: Task Summary and Real-Time Monitoring:
After a task begins running, users will receive a task summary notification via email.
Click the "View Results" button to access the real-time
monitoring page
for detailed progress tracking.
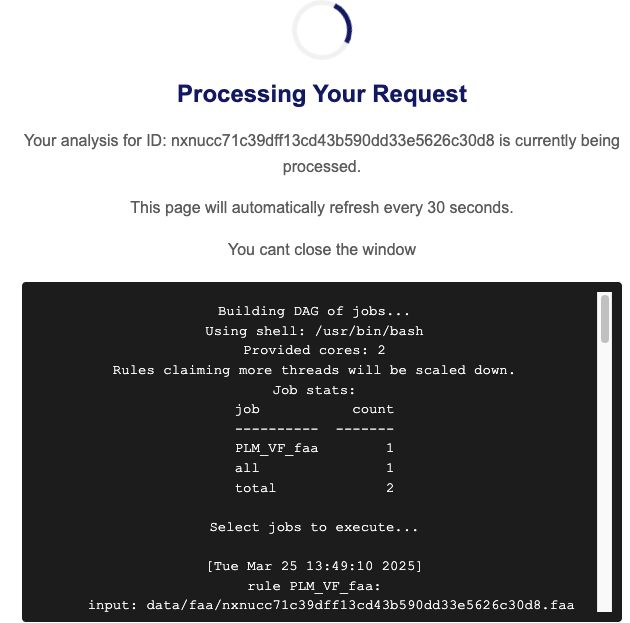
Step 4: Accessing Task Results:
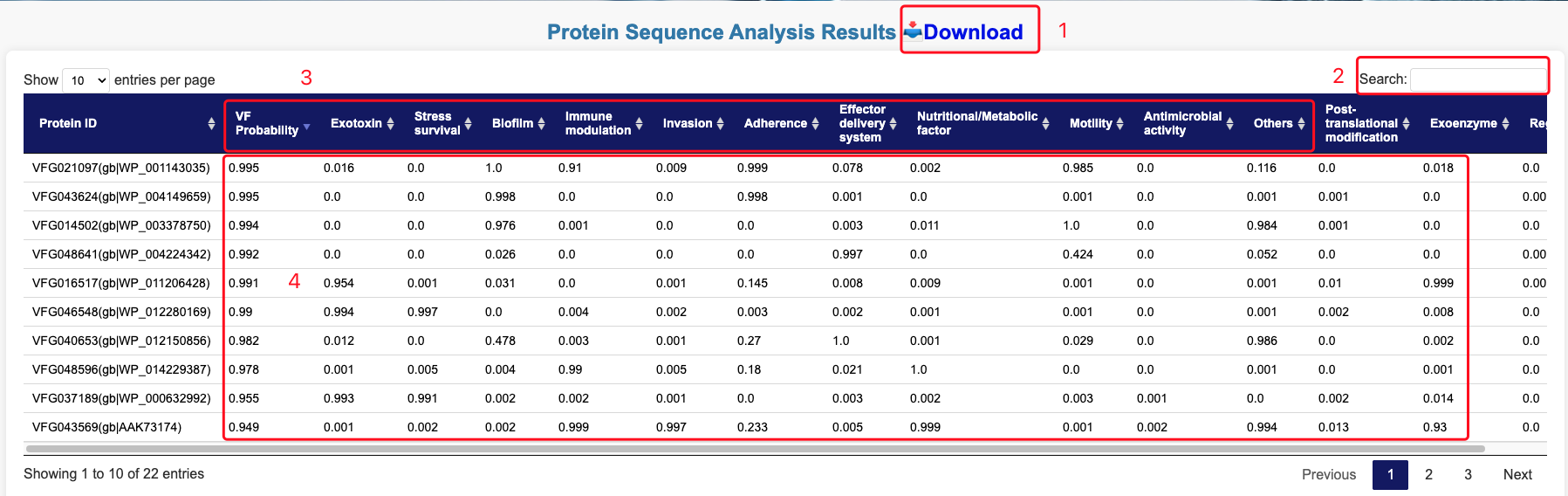
Once the task is completed, the system automatically redirects you to the results page.
- 1. Download the results in the CSV format.
- 2. Search/Filter: Refine or search for specific results directly within the page
- 3. Prediction Results Table: Sequence ID, Virulence Factor Category Name.
- 4. Probability of Virulence Factor Prediction (confidence score for virulence factor identification) and Virulence Factor Category Probability (confidence score for assigned category classification).
Source code:
Contact
If you have problems with the web server or submit a bug, you can contact Jian Ouyang (ouyangjian12@163.com).Please contact Dr. Chen (nxsfcc@nxnu.edu.cn) for comments on our website features or for adding new features or data.
Declaration of interest
This tool is for academic purposes and research use only. Any commercial use is subject for authorization from East China Nomral University (ECNU). Please contact us at tieliushi@yahoo.com.